Methods for plotting sp spatial objects with ggplot2.
Usage
# S3 method for class 'SpatialPoints'
gg(data, mapping = NULL, crs = NULL, ...)
# S3 method for class 'SpatialLines'
gg(data, mapping = NULL, crs = NULL, ...)
# S3 method for class 'SpatialPolygons'
gg(data, mapping = NULL, crs = NULL, ...)
# S3 method for class 'SpatialGridDataFrame'
gg(data, ...)
# S3 method for class 'SpatialPixelsDataFrame'
gg(data, mapping = NULL, crs = NULL, mask = NULL, ...)
# S3 method for class 'SpatialPixels'
gg(data, ...)Arguments
- data
A
Spatial*object.- mapping
Aesthetic mappings created by
aesused to update the default mapping. Uness specified otherwise below, the default mapping isggplot2::aes( x = .data[[sp::coordnames(data)[1]]], y = .data[[sp::coordnames(data)[2]]] )- crs
A
sp::CRSobject defining the coordinate system to project the data to before plotting.- ...
Arguments passed on to
geom_*.- mask
A
sp::SpatialPolygonsobject defining the region that is plotted.
Functions
gg(SpatialPoints): Geom forSpatialPointsobjects. This function coerces theSpatialPointsinto adata.frameand usesgeom_pointto plot the points. Requires theggplot2package.gg(SpatialLines): Geom for SpatialLines objects.Extracts start and end points of the lines and calls
geom_segmentto plot lines between them.mapping: Aesthetic mappings created byggplot2::aesorggplot2::aes_used to update the default mapping. The default mapping isggplot2::aes( x = .data[[sp::coordnames(data)[1]]], y = .data[[sp::coordnames(data)[2]]], xend = .data[[paste0("end.", sp::coordnames(data)[1])]], yend = .data[[paste0("end.", sp::coordnames(data)[2])]])gg(SpatialPolygons): Geom for SpatialPolygons objects. Uses theggplot2::fortify()function to turn theSpatialPolygonsobjects into adata.frame. Then callsgeom_polygonto plot the polygons.Unless specified by the user, the argument
alpha = 0.2(alpha level for polygon filling) is added.Up to version
2.10.0, theggpolypathpackage was used to ensure proper plotting for polygons, since theggplot2::geom_polygonfunction doesn't always handle geometries with holes properly. After2.10.0, the object is converted tosfformat and passed on togg.sf()instead, asggplot2version3.4.4deprecated the internally usedggplot2::fortify()method forSpatialPolygons/DataFrameobjects.gg(SpatialGridDataFrame): Geom for SpatialGridDataFrame objectsCoerces input
SpatialGridDataFrametoSpatialPixelsDataFrameand callsgg.SpatialPixelsDataFrame()to plot it.gg(SpatialPixelsDataFrame): Geom for SpatialPixelsDataFrame objects.Coerces
SpatialPixelsDataFrameinput todata.frameand usesgeom_tileto plot it.mapping: Aesthetic mappings created byaesused to update the default mapping. The default mapping isggplot2::aes( x = .data[[sp::coordnames(data)[1]]], y = .data[[sp::coordnames(data)[2]]], fill = .data[[names(data)[[1]]]] )gg(SpatialPixels): Geom for SpatialPixels objectsConverts the input to
SpatialPointsand calls [gg.SpatialPoints()` to plot it.
See also
Other geomes:
gg(),
gg.RasterLayer(),
gg.SpatRaster(),
gg.data.frame(),
gg.fm_mesh_1d(),
gg.fm_mesh_2d(),
gg.matrix(),
gg.sf()
Examples
# \donttest{
if (require("ggplot2", quietly = TRUE) &&
bru_safe_terra(quietly = TRUE) &&
bru_safe_sp() &&
require("sp")) {
# Load Gorilla data
gorillas <- inlabru::gorillas_sf
gcov <- gorillas_sf_gcov()
elev <- terra::as.data.frame(gcov$elevation, xy = TRUE)
elev <- sf::as_Spatial(sf::st_as_sf(elev, coords = c("x", "y")))
# Turn elevation covariate into SpatialGridDataFrame
elev <- sp::SpatialPixelsDataFrame(elev, data = as.data.frame(elev))
# Plot Gorilla elevation covariate provided as SpatialPixelsDataFrame.
# The same syntax applies to SpatialGridDataFrame objects.
ggplot() +
gg(elev)
# Add Gorilla survey boundary and nest sightings
ggplot() +
gg(elev) +
gg(gorillas$boundary, alpha = 0.0, col = "red") +
gg(gorillas$nests)
# Load pantropical dolphin data
mexdolphin <- inlabru::mexdolphin_sp()
# Plot the pantropical survey boundary, ship transects, and dolphin
# sightings
ggplot() +
gg(mexdolphin$ppoly) + # survey boundary as SpatialPolygon
gg(mexdolphin$samplers) + # ship transects as SpatialLines
gg(mexdolphin$points) # dolphin sightings as SpatialPoints
# Change color
ggplot() +
gg(mexdolphin$ppoly, color = "green") + # survey boundary; SpatialPolygon
gg(mexdolphin$samplers, color = "red") + # ship transects; SpatialLines
gg(mexdolphin$points, color = "blue") # dolphin sightings; SpatialPoints
# Visualize data annotations: line width by segment number
names(mexdolphin$samplers) # 'seg' holds the segment number
ggplot() +
gg(mexdolphin$samplers, aes(color = seg))
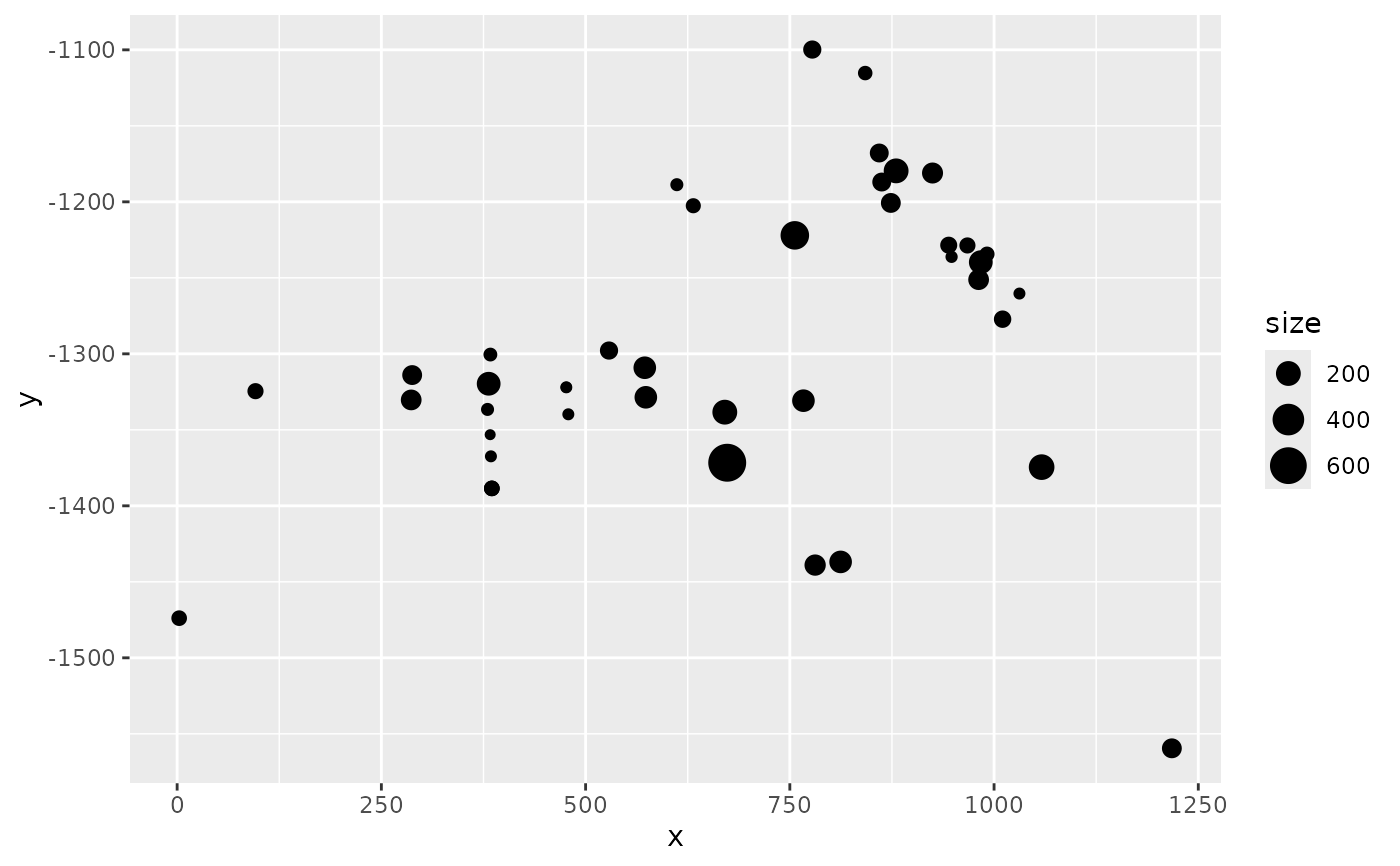
# Visualize data annotations: point size by dolphin group size
names(mexdolphin$points) # 'size' holds the group size
ggplot() +
gg(mexdolphin$points, aes(size = size))
}
#> Loading required package: sp
 # }
if (require("ggplot2", quietly = TRUE) &&
bru_safe_terra(quietly = TRUE) &&
bru_safe_sp()) {
# Load Gorilla data
gcov <- gorillas_sf_gcov()
elev <- terra::as.data.frame(gcov$elevation, xy = TRUE)
pxl <- sf::as_Spatial(sf::st_as_sf(elev, coords = c("x", "y")))
# Turn elevation covariate into SpatialPixels
pxl <- sp::SpatialPixels(pxl)
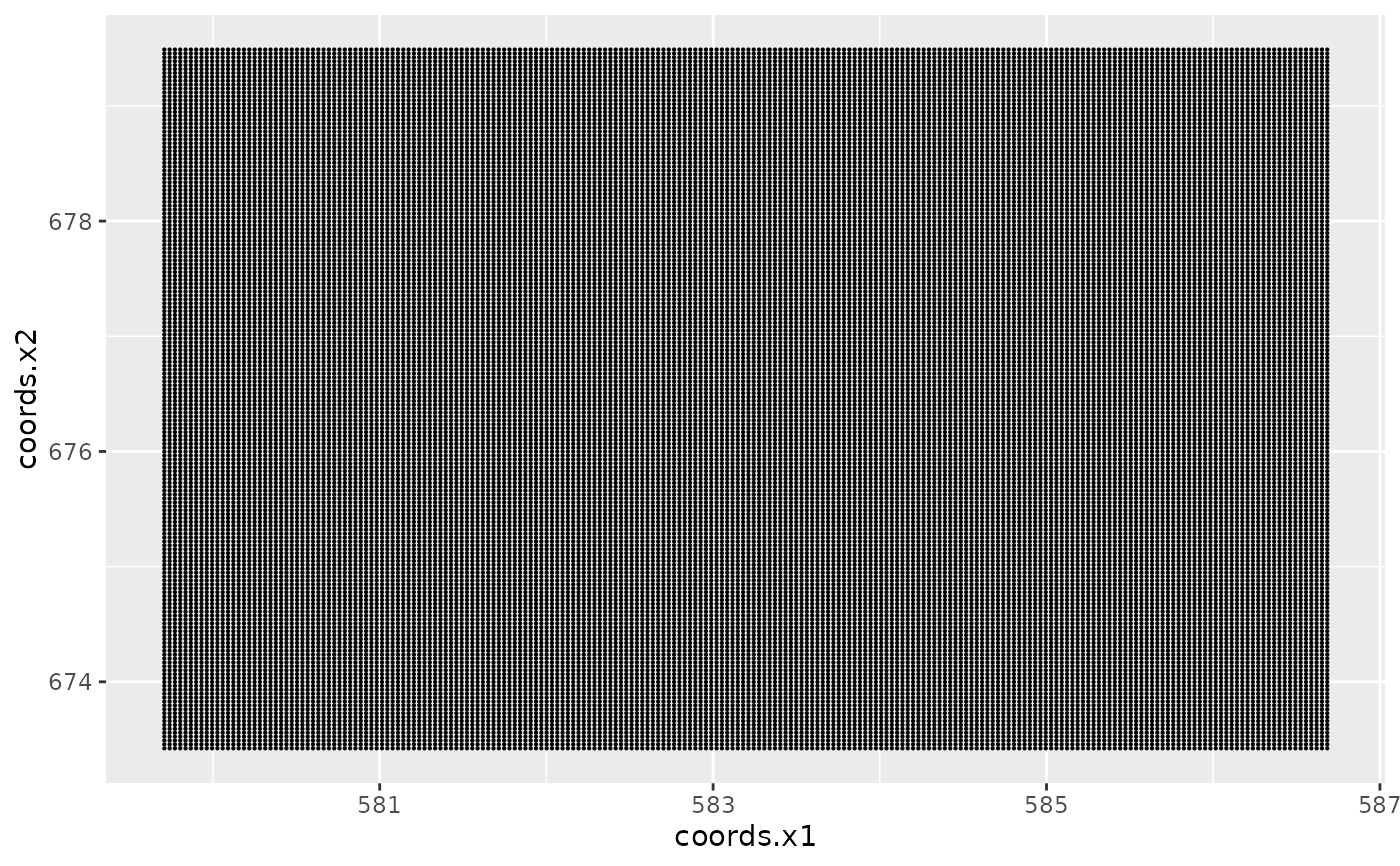
# Plot the pixel centers
ggplot() +
gg(pxl, size = 0.1)
}
# }
if (require("ggplot2", quietly = TRUE) &&
bru_safe_terra(quietly = TRUE) &&
bru_safe_sp()) {
# Load Gorilla data
gcov <- gorillas_sf_gcov()
elev <- terra::as.data.frame(gcov$elevation, xy = TRUE)
pxl <- sf::as_Spatial(sf::st_as_sf(elev, coords = c("x", "y")))
# Turn elevation covariate into SpatialPixels
pxl <- sp::SpatialPixels(pxl)
# Plot the pixel centers
ggplot() +
gg(pxl, size = 0.1)
}