Random Fields in 2D
Finn Lindgren
Generated on 2026-05-22
Source:vignettes/articles/random_fields_2d.Rmd
random_fields_2d.RmdSetting things up
Make a shortcut to a nicer colour scale:
colsc <- function(...) {
scale_fill_gradientn(
colours = rev(RColorBrewer::brewer.pal(11, "RdYlBu")),
limits = range(..., na.rm = TRUE)
)
}Modelling on 2D domains
We will now construct a 2D model, generate a sample of a random field, and attempt to recover the field from observations at a few locations. Tomorrow, we will look into more general mesh constructions that adapt to irregular domains.
First, we build a high resolution mesh for the true field, using low level INLA functions
bnd <- spoly(
data.frame(
easting = c(0, 10, 10, 0),
northing = c(0, 0, 10, 10)
),
format = "sf"
)
## For fmesher 0.3.0:
## mesh_fine <- fm_mesh_2d_inla(boundary = bnd, max.edge = 0.2)
## For fmesher >= 0.4.0:
mesh_fine <- fm_mesh_2d(
loc = fm_hexagon_lattice(bnd, edge_len = 0.2),
boundary = bnd,
max.edge = 0.3
)
ggplot() +
geom_fm(data = mesh_fine)
# Note: the priors here will not be used in estimation
matern_fine <-
inla.spde2.pcmatern(mesh_fine,
prior.sigma = c(1, 0.01),
prior.range = c(1, 0.01)
)
true_range <- 4
true_sigma <- 1
true_Q <- inla.spde.precision(matern_fine,
theta = log(c(true_range, true_sigma))
)What is the pointwise standard deviation of the field? Along straight boundaries, the variance is twice the target variance. At corners the variance is 4 times as large.
true_sd <- diag(inla.qinv(true_Q))^0.5
ggplot() +
gg(mesh_fine, color = true_sd) +
coord_equal()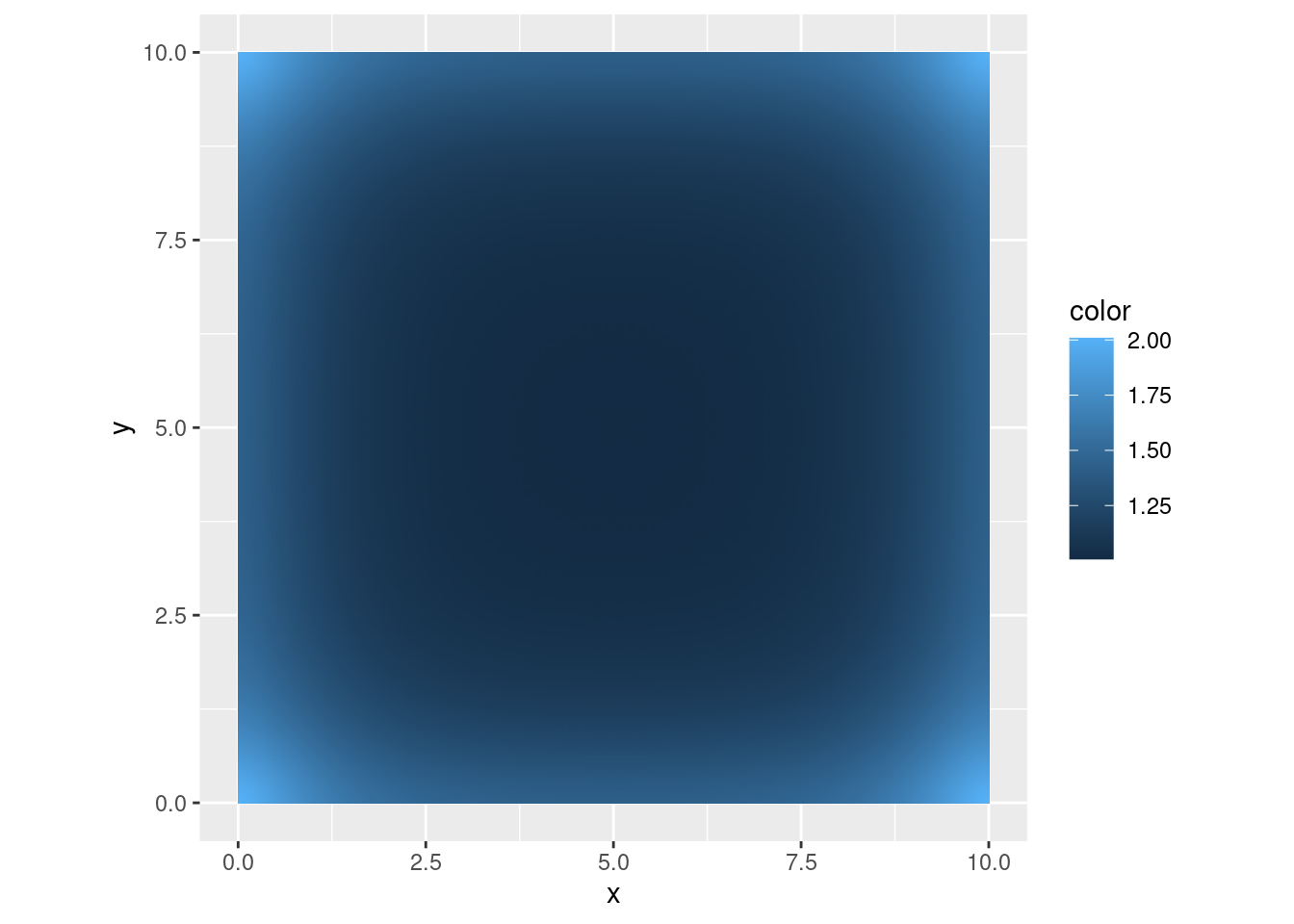
Generate a sample from the model:
true_field <- inla.qsample(1, true_Q)[, 1]
truth <- expand.grid(
easting = seq(0, 10, length = 100),
northing = seq(0, 10, length = 100)
)
truth <- sf::st_as_sf(truth, coords = c("easting", "northing"))
truth$field <- fm_evaluate(
mesh_fine,
loc = truth,
field = true_field
)
pl_truth <- ggplot() +
gg(truth, aes(fill = field), geom = "tile") +
ggtitle("True field")
## Or with another colour scale:
csc <- colsc(truth$field)
(pl_truth | (pl_truth + csc))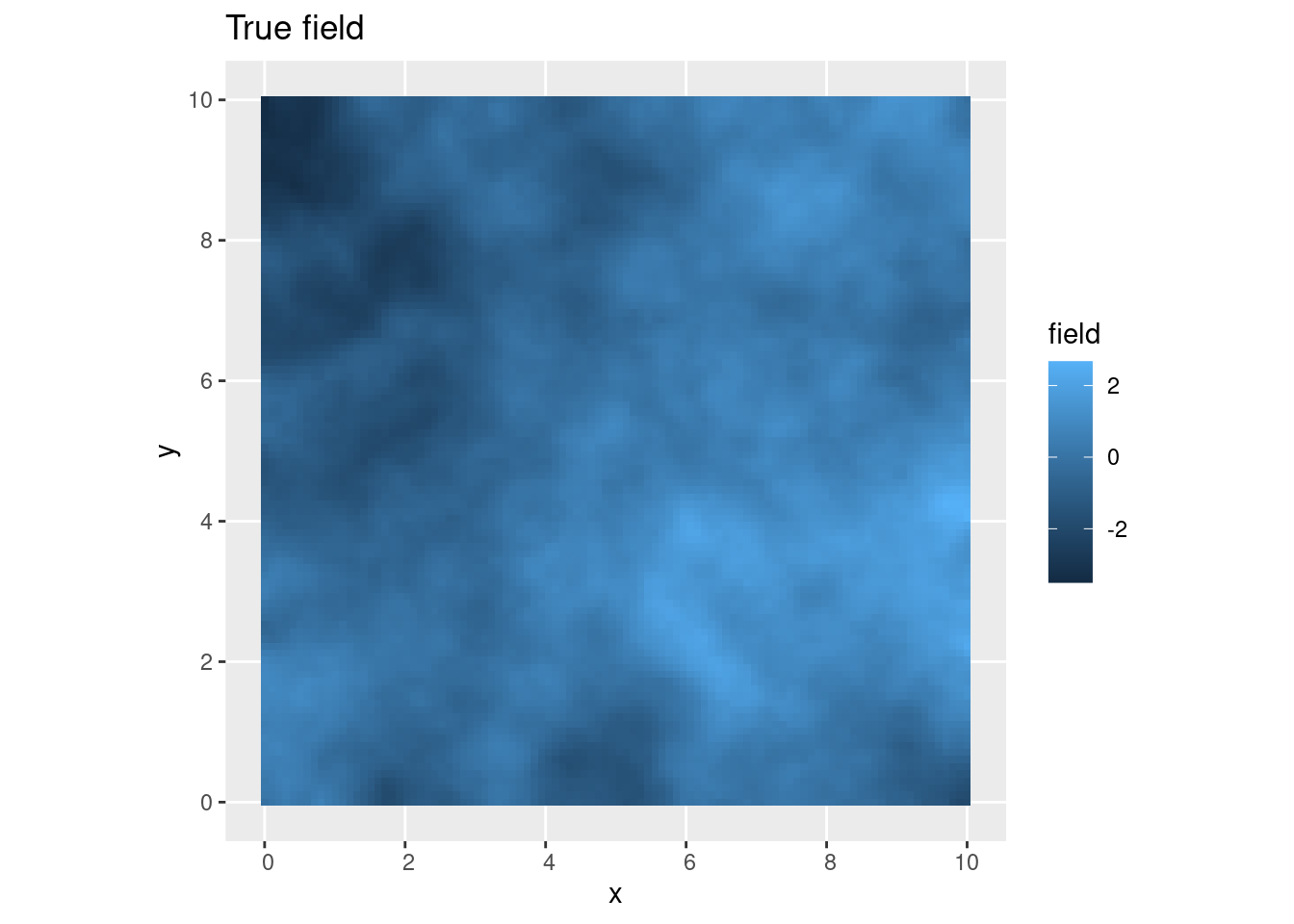
Extract observations from some random locations:
n <- 200
mydata <- sf::st_as_sf(
data.frame(easting = runif(n, 0, 10), northing = runif(n, 0, 10)),
coords = c("easting", "northing")
)
mydata$observed <-
fm_evaluate(
mesh_fine,
loc = mydata,
field = true_field
) +
rnorm(n, sd = 0.4)
ggplot() +
gg(mydata, aes(col = observed))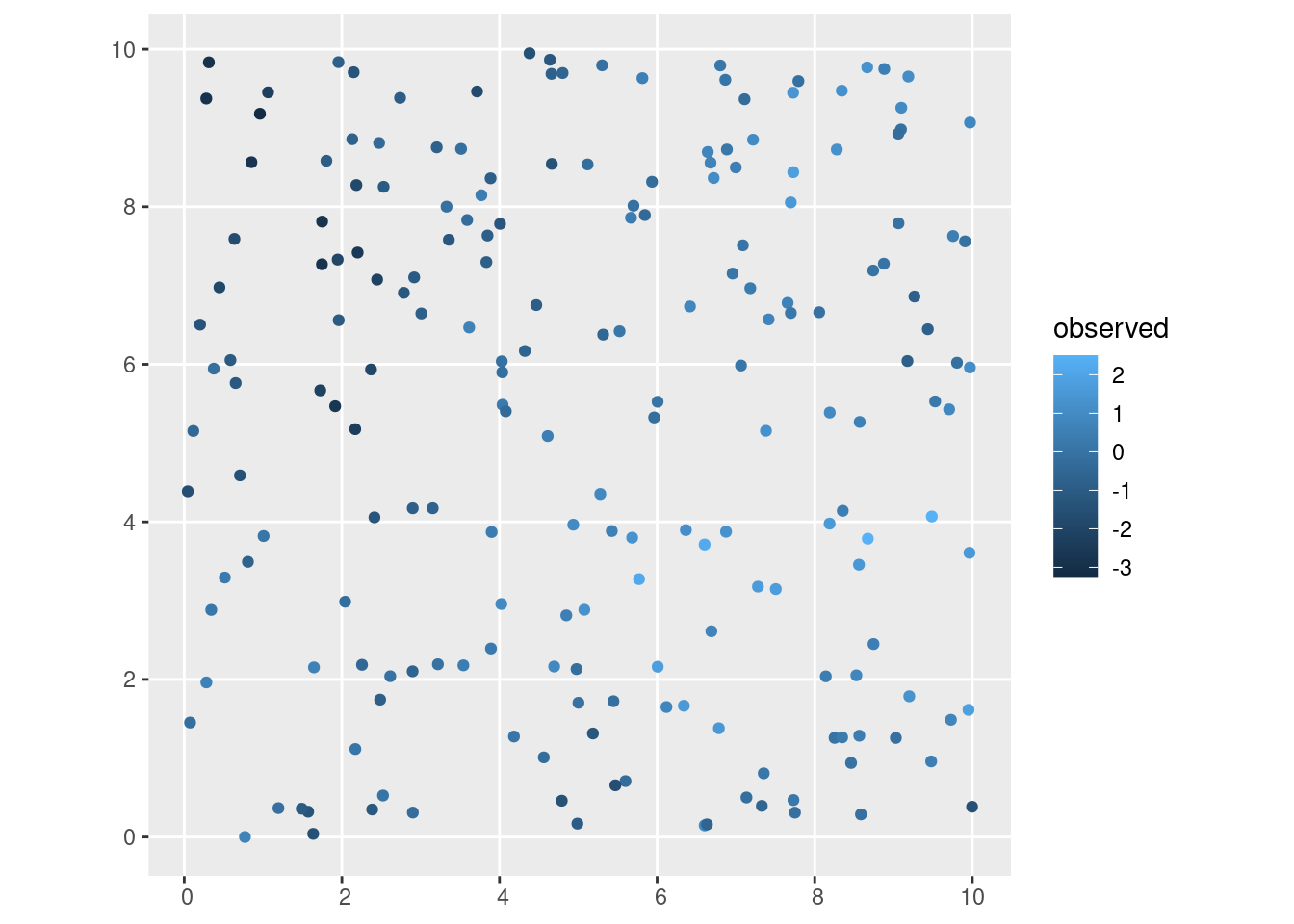
Estimating the field
Construct a mesh covering the data:
## For fmesher 0.3.0:
## mesh <- fm_mesh_2d_inla(
## boundary = bnd,
## max.edge = 0.5
## )
## For fmesher >= 0.4.0
mesh <- fm_mesh_2d(
loc = fm_hexagon_lattice(bnd, edge_len = 0.5),
boundary = bnd,
max.edge = 0.6
)
ggplot() +
geom_fm(data = mesh)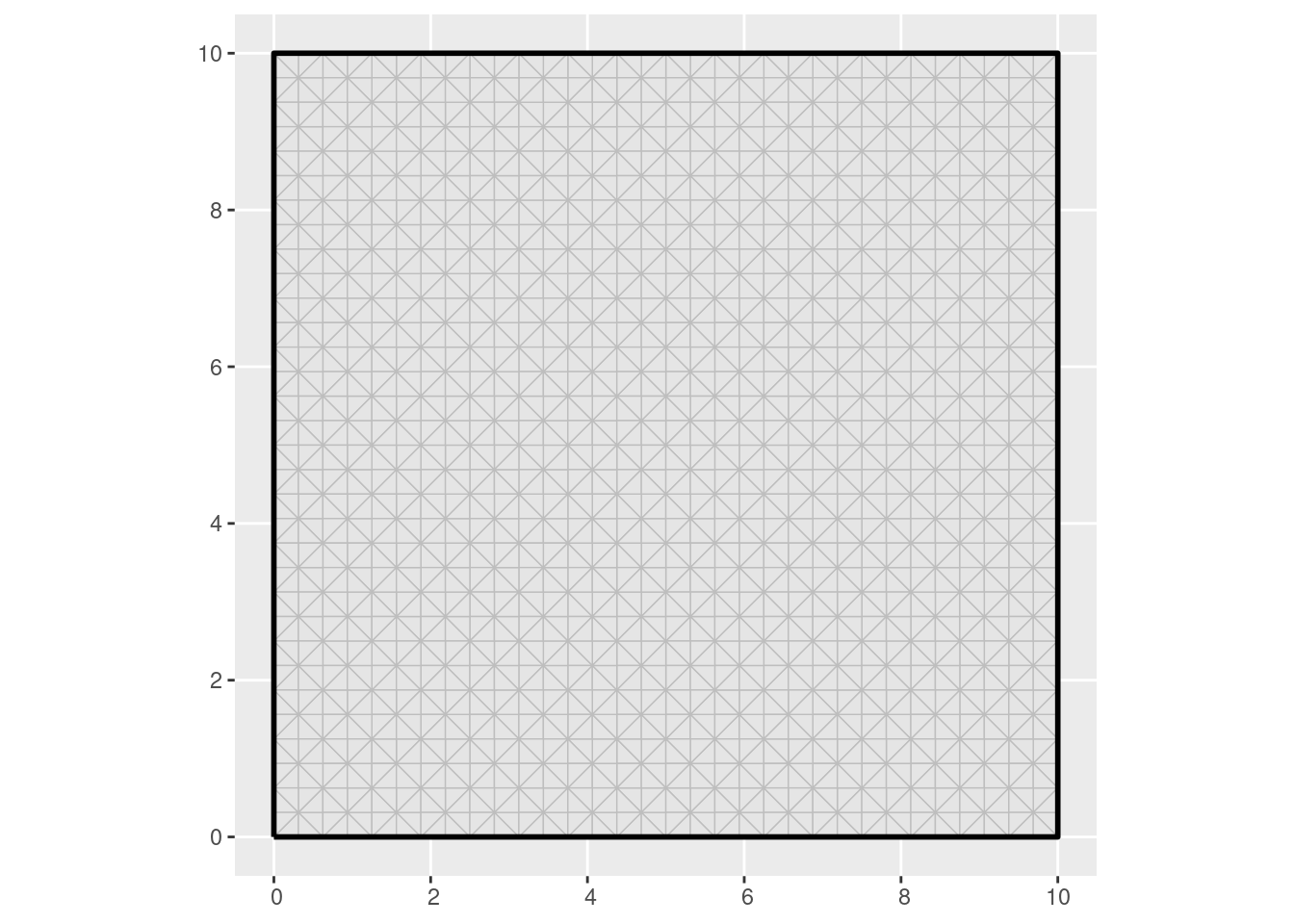
Construct an SPDE model object for a Matern model:
matern <-
inla.spde2.pcmatern(mesh,
prior.sigma = c(10, 0.01),
prior.range = c(1, 0.01)
)Specify the model components:
cmp <- observed ~ field(geometry, model = matern) + Intercept(1)Fit the model and inspect the results:
Predict the field on a lattice, and generate a single realisation from the posterior distribution:
pix <- fm_pixels(mesh, dims = c(200, 200))
pred <- predict(
fit,
pix,
~ field + Intercept
)
samp <- generate(
fit,
pix,
~ field + Intercept,
n.samples = 1
)
pred$sample <- samp[, 1]Compare the truth to the estimated field (posterior mean and a sample from the posterior distribution):
pl_posterior_mean <- ggplot() +
gg(pred, geom = "tile") +
gg(bnd, alpha = 0) +
ggtitle("Posterior mean")
pl_posterior_sample <- ggplot() +
gg(pred, aes(fill = sample), geom = "tile") +
gg(bnd, alpha = 0) +
ggtitle("Posterior sample")
# Common colour scale for the truth and estimate:
csc <- colsc(truth$field, pred$mean, pred$sample)
((pl_truth + csc) |
(pl_posterior_mean + csc) |
(pl_posterior_sample + csc))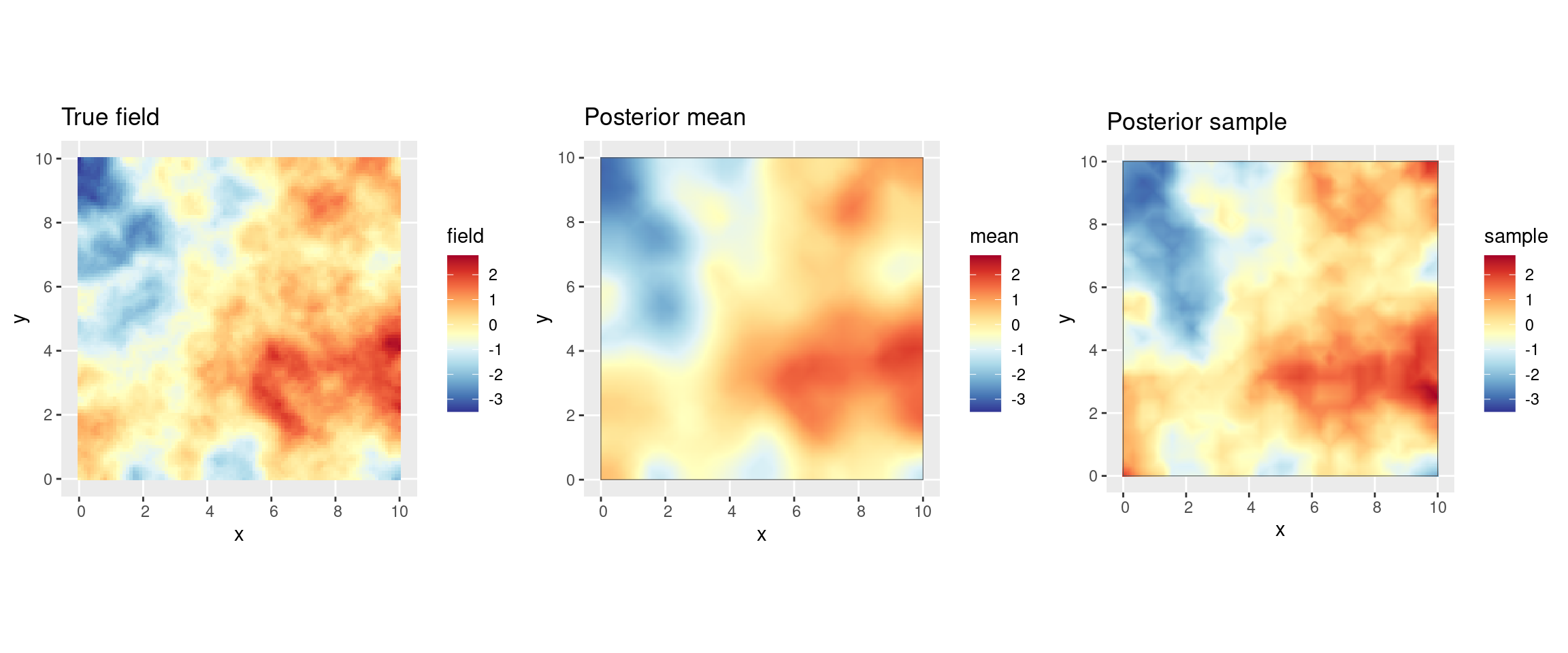
Plot the SPDE parameter and fixed effect parameter posteriors.
int.plot <- plot(fit, "Intercept")
spde.range <- spde.posterior(fit, "field", what = "range")
spde.logvar <- spde.posterior(fit, "field", what = "log.variance")
range.plot <- plot(spde.range)
var.plot <- plot(spde.logvar)
(range.plot / var.plot / int.plot)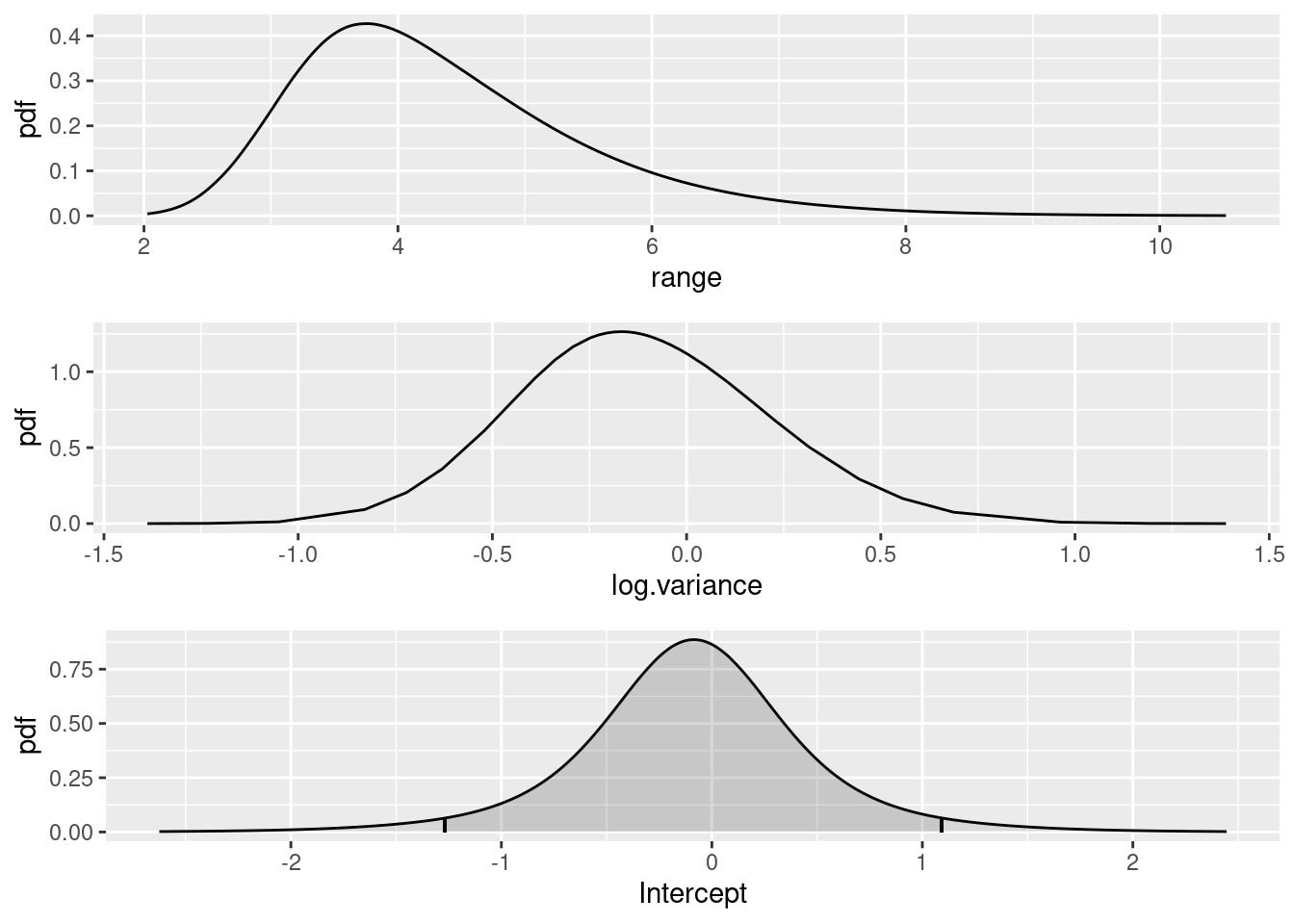
Look at the correlation function if you want to:
corplot <- plot(spde.posterior(fit, "field", what = "matern.correlation"))
covplot <- plot(spde.posterior(fit, "field", what = "matern.covariance"))
(covplot / corplot)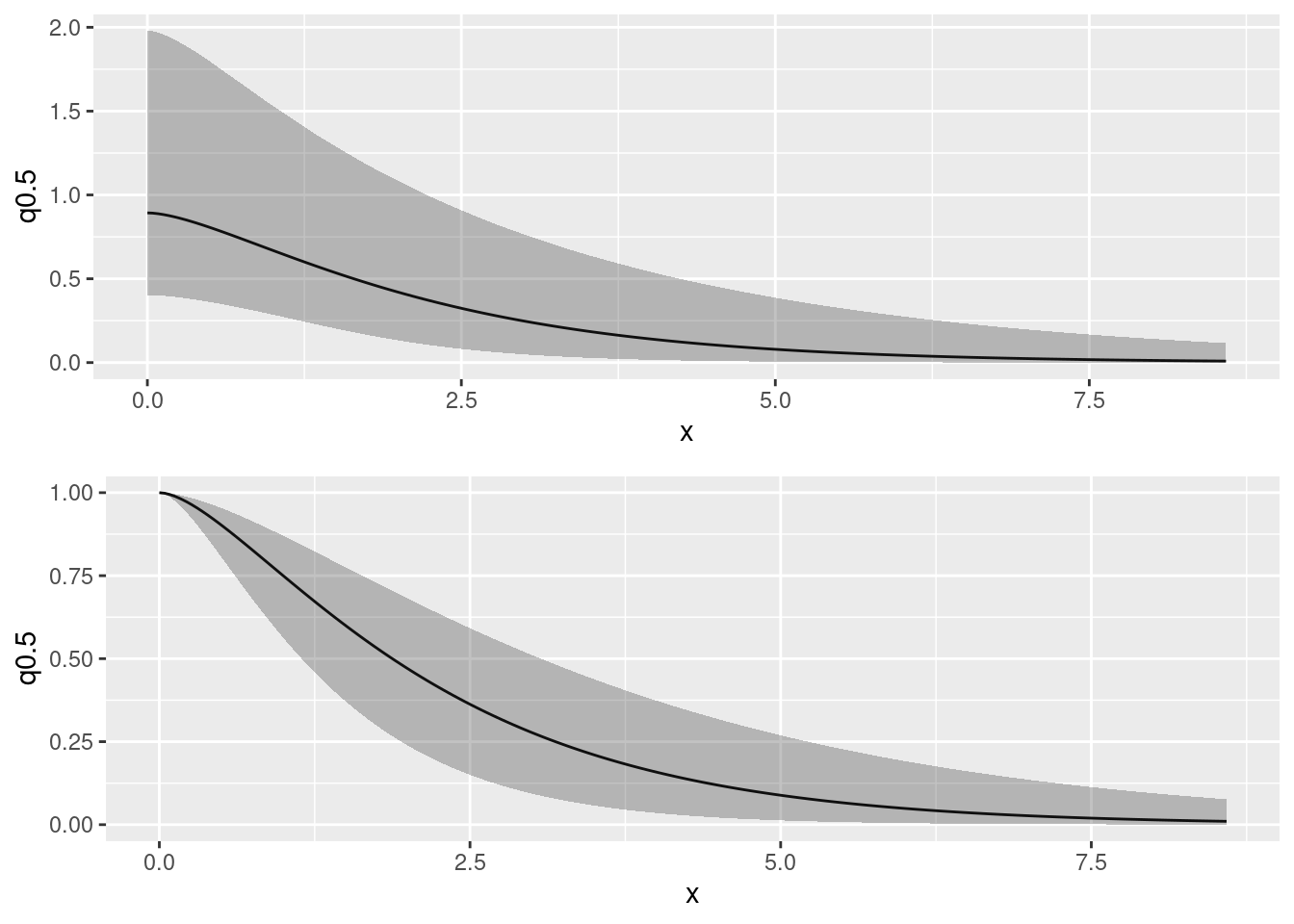
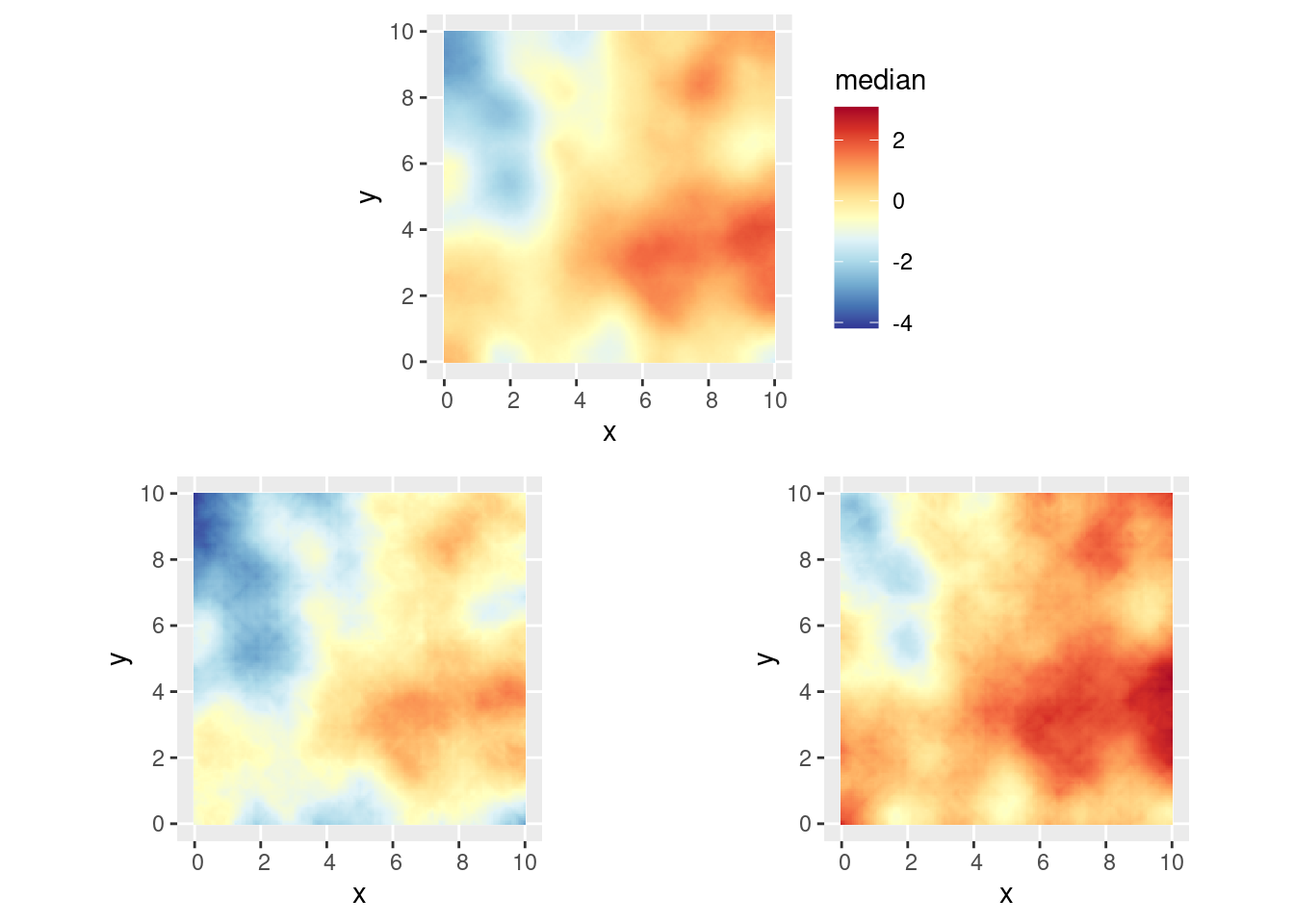
You can plot the median, lower 95% and upper 95% density surfaces as
follows (assuming that the predicted intensity is in object
pred).
csc <- colsc(
pred[["median"]],
pred[["q0.025"]],
pred[["q0.975"]]
) ## Common colour scale from SpatialPixelsDataFrame
gmedian <- ggplot() +
gg(pred["median"], geom = "tile") +
csc
glower95 <- ggplot() +
gg(pred["q0.025"], geom = "tile") +
csc +
theme(legend.position = "none")
gupper95 <- ggplot() +
gg(pred["q0.975"], geom = "tile") +
csc +
theme(legend.position = "none")
(gmedian / (glower95 | gupper95))